Lotka-Volterra 2D¶
The Original Model in Ordinary Differential Equations¶
[1]:
%matplotlib inline
from ecell4 import *
[2]:
alpha = 1
with reaction_rules():
~u > u | u * (1 - v)
~v > v | alpha * v * (u - 1)
m = get_model()
[3]:
run_simulation(15, {'u': 1.25, 'v': 0.66}, model=m)
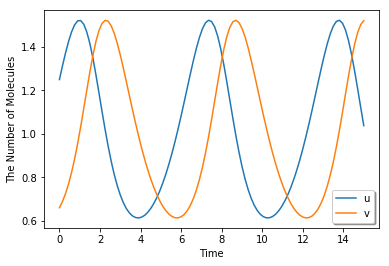
The Modified Model Decomposed into Elementary Reactions¶
[4]:
alpha = 1
with species_attributes():
u | {'D': 0.1}
v | {'D': 0.1}
with reaction_rules():
u > u + u | 1.0
u + v > v | 1.0
u + v > u + v2 | alpha
v2 > v + v | alpha * 10000.0
v > ~v | alpha
m = get_model()
[5]:
run_simulation(40, {'u': 1.25 * 1600, 'v': 0.66 * 1600}, volume=1600, model=m)
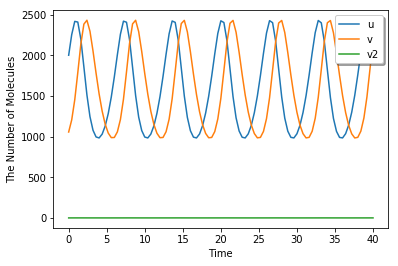
[6]:
run_simulation(40, {'u': 1.25 * 1600, 'v': 0.66 * 1600}, volume=1600, model=m, solver='gillespie')
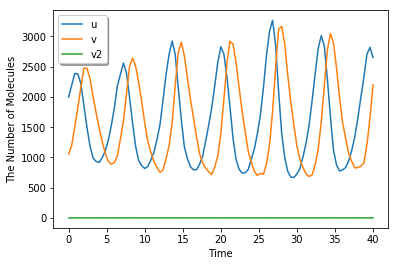
[7]:
alpha = 1
with species_attributes():
u | v | {'D': 0.1}
with reaction_rules():
u > u + u | 1.0
u + v > v | 1.0
u + v > u + v + v | alpha
v > ~v | alpha
m = get_model()
[8]:
run_simulation(40, {'u': 1.25 * 1600, 'v': 0.66 * 1600}, volume=1600, model=m)
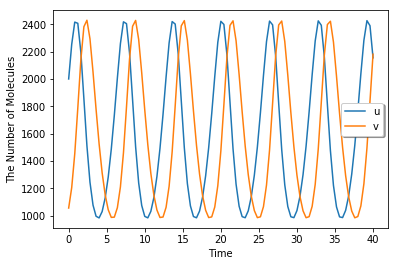
[9]:
run_simulation(40, {'u': 1.25 * 1600, 'v': 0.66 * 1600}, volume=1600, model=m, solver='gillespie')
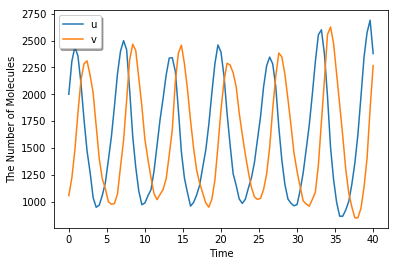
A Lotka-Volterra-like Model in 2D¶
[10]:
from ecell4_base.core import *
from ecell4_base import meso
[11]:
rng = GSLRandomNumberGenerator()
rng.seed(0)
[12]:
w = meso.World(Real3(40, 40, 1), Integer3(160, 160, 1), rng)
w.bind_to(m)
[13]:
V = w.volume()
print(V)
1600.0
[14]:
w.add_molecules(Species("u"), int(1.25 * V))
w.add_molecules(Species("v"), int(0.66 * V))
[15]:
sim = meso.Simulator(w)
obs1 = FixedIntervalNumberObserver(0.1, ('u', 'v', 'v2'))
obs2 = FixedIntervalHDF5Observer(2, "test%03d.h5")
[16]:
sim.run(100, (obs1, obs2))
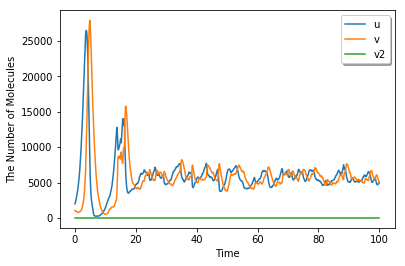
[17]:
viz.plot_number_observer(obs1)
[18]:
viz.plot_world(w, radius=0.2)
[19]:
viz.plot_movie_with_attractive_mpl(
obs2, linewidth=0, noaxis=True, figsize=6, whratio=1.4,
angle=(-90, 90, 6), bitrate='10M')
[20]:
# from zipfile import ZipFile
# with ZipFile('test.zip', 'w') as myzip:
# for i in range(obs2.num_steps()):
# myzip.write('test{:03d}.h5'.format(i))